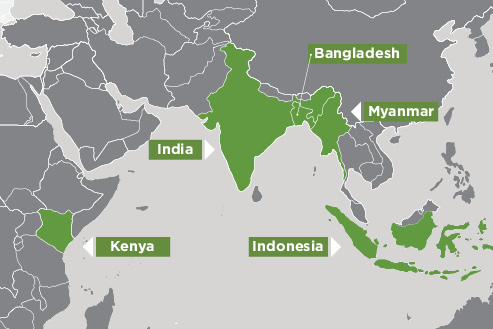
Overview
This project aims to increase farmer income, systems productivity and food security by delivering high-yielding, disease and pest resistant mungbean varieties to smallholder farmers.
Declining crop yields, increased agricultural risks, diminishing soil fertility and environmental degradation are some of the main challenges which continue to threaten societal goals of improving food, income and nutrition security especially in smallholder farming.
There is large demand for options for sustainable intensification of tropical cropping systems of Africa, South Asia and Australia.
The project will build on current ACIAR projects researching mungbean agronomy and contribution to cropping system intensification in Myanmar and Bangladesh.
Expected project outcomes
- Creating New traits and methods for mungbean improvement.
- Increasing knowledge on legumes and legumes traits.
- Implementing capacity development of breeders including training on better database management, application of marker assisted selection, statistical methods in breeding and adoption of integrated pest management strategies.
- Gaining access to new traits introgressed in breeding lines, allowing them to deliver new varieties, more adapted to farmers’ needs, faster.
Summary of outcomes to date
2021–22
- The annual meeting of the network was conducted online on 28 June 2022. Discussions were held on the progress on phenotyping, genotyping and on the testing of the newly developed lines. Sequencing efforts (Illumina) of mungbean accessions of the WorldVeg mini-core, the Australian diversity panel and segregating populations yielded up to now 2 terabasepairs of sequence information that is currently used to develop a mungbean pangenome and map traits of interest for breeding. A pilot experiment on a selected germplasm set revealed the optimal planting density for automatized high throughput phenotyping of mungbean plant architecture.
- In Australia, the entire NAM population was sown and evaluated for key flowering traits. Data collected from this trial will enable a genome-wide associated study to be dissect genomic regions associated with these traits.
- The further screening of the mini-core accessions/breeding lines has revealed potential sources of resistance to anthracnose, powdery mildew, Fusarium wilt, and cowpea aphid, thrips and stemfly.
- A set of 42 accessions from 13 Vigna species endemic to India was screened against three bruchid species (Callosobruchus. chinensis L., C. maculatus F. and C. analis L., which led to the identification of accessions with resistance.
- The project partners are actively engaged in the introgression of traits from donors identified during IMIN1 and the testing of the high performing lines is in progress.